After a long pandemic break, PyCon DE & PyData is back again! I attended previous conferences remotely, and this year was the first time when I arrived to Berlin to be there in person. The conference was great, with plenty of amazing talks and insights from leading Python engineers and data scientists. And while the memories are still fresh, I would like to structure the collected notes and briefly summarize attended presentations.
 The logo of PyCon DE & PyData Berlin 2022
The logo of PyCon DE & PyData Berlin 2022
Many of the talks are accompanied by example repositories, slides, and more elaborate blog posts for in-depth learning. I include these links here when available. Also, it was a so-called hybrid conference with both remote and in-person participants. Each of the talks was recorded and will be available online after a while.
Before we start, a small tip for people who never attended the conference. As you can guess from its title, it covers two major groups of topics. The first one is about software engineering in Python in general. And the second one encompasses talks about doing data analysis and machine learning using Python’s numerical stack. (Which is probably not a big surprise for people closely following the language’s development and applications.) Therefore, the topics described here can be roughly categorized into two (overlapping) sets: 🐍 Python Engineering and ⚛️ Data Science️. I mark each summary with these emojis to make it a bit easier to navigate.
Post contents
🐍 Adding a new operator to Python
In his talk Demystifying Python’s Internals: Diving into CPython by implementing a pipe operator, Sebastiaan Zeeff showed us how to bring a completely new operator into Python language. That sounds like something super-difficult and requiring many years of experience in developing compilers and language parsers. However, in reality, it turned out to be not that difficult! That’s why I liked this talk so much. It strives to reach the goal that looks quite intimidating at first glance: changing the grammar of Python interpreter. And then the speaker disassembles it into smaller steps that are (relatively) easy to implement.
The speaker forked the official CPython implementation and changed it to enable function pipes which you many know about from functional programming languages:
def double(x: float) -> float:
return x * x
def increment(x: float) -> float:
return x + 1
result = 2 |> double |> increment |> double # 10
Isn’t that great? Also, the speaker points out to two valuable sources of information for people who want to know more about CPython internals and/or get involved into the language development process. This talk ignited my interest in hacking CPython to get more insights into its implementation and get a better understanding of how the language works.
🐍 ⚛️ Making Python Fast(er)
Stefan Behnel presented a talk Fast native data structures: C/C++ from Python where he shares insights about using Cython to enable fast C modules that speed up the execution of Python code.
The talk includes several examples, from the most basic ones to more involved. The easiest way to get some performance gains is to annotate your variables with Cython data types as the following snippet shows.
import cython
def sum_of_int_squares(ints: cython.long[:]):
i: cython.long
s: cython.long = 0
for i in ints:
s += i * i
return s
Note that during his talk, the speaker used Jupyter and magic commands to make the presentation interactive and quick. However, if you want to enable Cython in your scripts, you’ll need an extra compilation step. Keep this in mind if decide to introduce it into your codebase.
You may wonder why to bring an extra compilation complexity if there are libraries like numpy and pandas that already rely on
C libraries to make computations fast. But Cython can bring a noticeable performance gains even in this case.
In his presentation, the speaker shows a more complex example where he uses C++ <algorithm.cpp> functions imported into Python
code to efficiently calculate a rolling median. The shown implementation works considerably (~25 times!) faster
compared to the computation time required when using np.median function.
⚛️ Copy-on-Write for Pandas Data Frames?
If you’re a pandas user, then the chances are high that sooner or later you will stumble upon the following warning:
SettingWithCopyWarning:
A value is trying to be set on a copy of a slice from a DataFrame.
Try using .loc[row_indexer,col_indexer] = value instead
See the caveats in the documentation: [...]
In his talk On Blocks, Copies and Views: updating pandas’ internals
Joris Van den Bossche reminds listeners how tricky the pandas memory management can be. It is not always clear
if you work on a data frame’s view or its copy. Which at least pushes developers into “defensive copying” all
around the codebase, like putting df = df.copy() before transforming the data in order to not modify it accidentally.
And at most produces subtle bugs and inefficiency.
Could we do better? Yes! The speaker, a pandas core contributor, talks about a possible solution that can be expressed using a single rule:
AnyDataFrameorSeriesderived from another in any way (e.g. with an indexing operation) always behaves as a copy.
It means that, for example, whenever a data frame subset is taken, it behaves as a new object and if modified, the original data frame doesn’t change.
Note the emphasis on the word “behaves” in the quote above. In case if we make a copy of a data frame each time when
a range of rows or columns is selected, it would be a very inefficient memory usage. Instead, a
copy-on-write (CoW) pattern should be implemented as a core data frame
feature to make sure that memory is allocated only when needed. Then we have the best of both worlds: derived data
frames don’t change the original one, but new memory is allocated only if a change happens. The “defensive copying”
isn’t needed and no more SettingWithCopy warnings raised.
The biggest challenge however is backwards incompatibility of this feature with previous versions of the library. For example, the following line of code will have a different behavior.
# Works, but will not if CoW is enabled
df["C"][df["A"] > 1] = 10
Currently, it mutates the df data frame, but with CoW enabled, it creates a copy which is immediately discarded.
temp = df["C"]
temp[df["A"] > 1] = 10
del temp
Of course, it would break all code that uses this syntax to modify data frames. Therefore, the discussion is still
going on. Nevertheless, this change should significantly improve pandas API and I believe that many people
(including myself) would like to see it in the library.
⚛️ Target Variable Transformation Pitfalls
Let’s say you have a non-negative right-skewed target variable, i.e., the number of sales in a store. You want to fit a regression model and use Root Mean-Squared Percentage Error (RMSPE) to assess the quality of predictions. So why not transform the target using the logarithm to make it more normally distributed? You do the transformation, and after model fitting, you transform the predicted target back, and compute the metric to estimate model’s quality. Sounds like a good idea, right?
Well, there is a problem. In his excellent talk Honey, I shrunk the target variable! Florian Wilhelm shows that by doing that, we essentially change the optimization problem. We’re not approximating the mean anymore, as we would expect, but rather for a median value. And as you can guess, it has a measurable impact on the quality of our forecast. In order to fix the issue, we should add a correction factor before transforming our prediction back from the log space. The exact value of correction depends on chosen metric.
My description here is intentionally brief, as, in addition to his talk, the speaker wrote a great blog post describing the problem very thoroughly. It also includes elaborated mathematical proofs showing why the correction is needed and what it means to approximate the median instead of the mean. As a kind of teaser to invite you to read the post, below you see a table taken from there, showing the percentage of improvement for each metric depending on the type of correction. (The lower value, the better.) Note that the impact on RMSPE, which is highlighted, is especially significant.
| RMSE | MAE | MAPE | RMSPE | |||||
|---|---|---|---|---|---|---|---|---|
| target | mean | std | mean | std | mean | std | mean | std |
| log & no corr | +3.42% | ±1.07%p | -0.09% | ±0.61%p | -10.99% | ±0.65% | -12.08% | ±4.14%p |
| log & sigma2 corr | +4.14% | ±0.84%p | -0.09% | ±0.61%p | -11.03% | ±0.74%p | -28.24% | ±3.35%p |
| log & fitted corr | +2.75% | ±1.00%p | -0.19% | ±0.58%p | -13.23% | ±0.68%p | -47.27% | ±5.37%p |
I highly recommend you to read through the post, as it gives very deep insights into the problem. For a summary: choose your transformations carefully! And make sure that the transformation is consistent with the metric you are optimizing.
⚛️ Visualizing Big Data
Python provides a great variety of visualization tools. In many cases, it is perfectly fine to use well-known and
widely adopted tools like matplotlib, or plotting functions attached to pandas. In case if some level interactivity is
needed, you can harness web rendering tools like plotly or altair. But what if your dataset is big?
I mean, not just a few thousand rows, but a million? Or even a billion? The browser is not able to handle such a
dataset and will crash. And even if it could, it would be impossible to distinguish between individual data points.
Fortunately, there are tools that can help you visualize big data, while keeping the good level of interactivity. The talk
Seeing the Needle and the Haystack by Jean-Luc Stevens sheds light on tools
for plotting huge datasets while keeping the plots interactive and responsive. It introduces an easy-to-use hvPlot API
that leverages HoloViews, Datashader, Bokeh, and Panel to build dashboards that do server-side rendering of
billions of data points without losing the ability to interactively inspect individual samples in the browser.
The speaker illustrates hvPlot capabilities by rendering huge (hundreds of millions) lat/long dataset in a Jupyter notebook,
highlighting a combination of performance and interactivity. The samples are aggregated depending on a zoom level. Each
data point keeps track of its meta-information and can be easily distinguished from others. (See a small example below.)
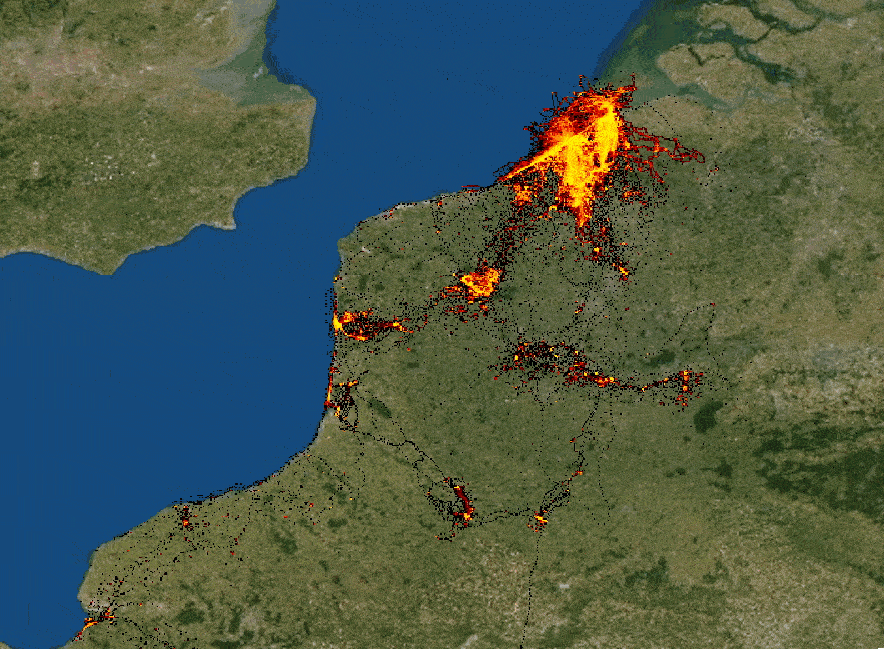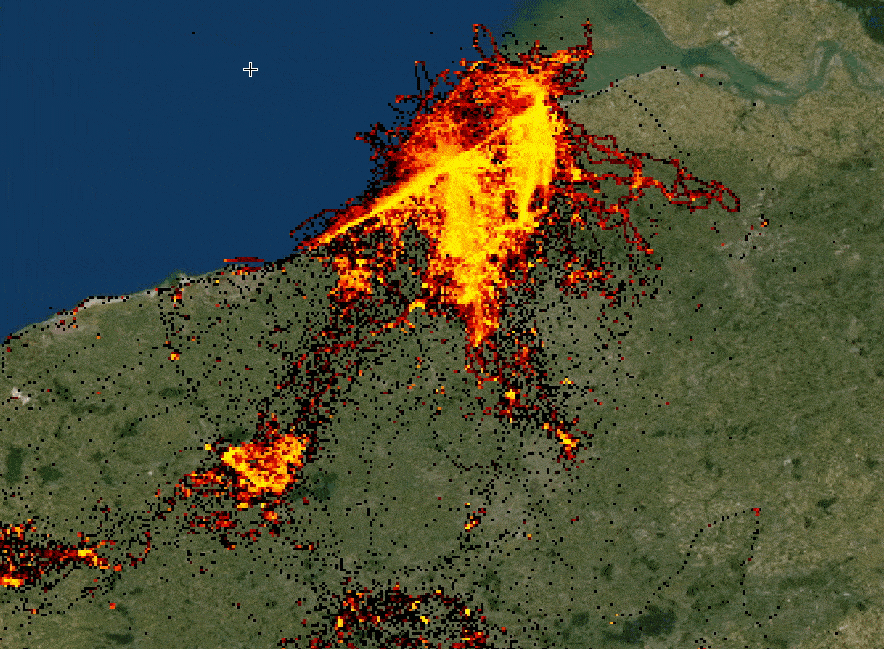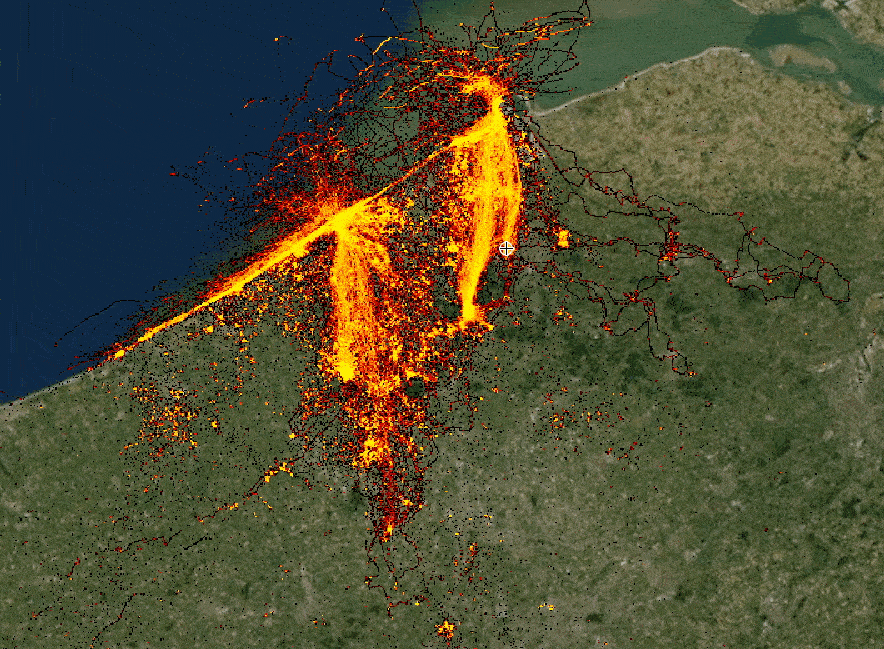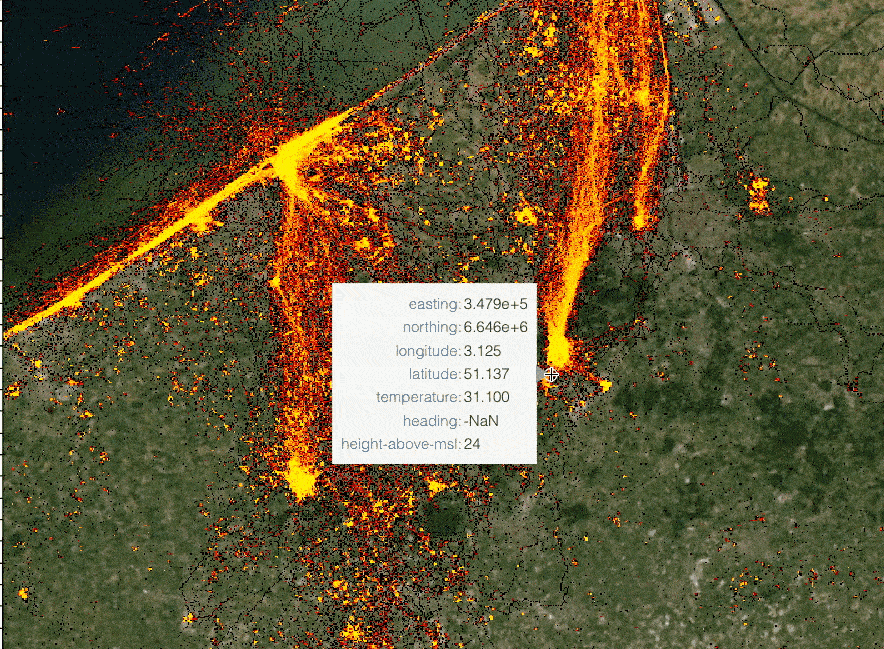 Interactive rendering of a huge dataset with data re-aggregation depending on zoom level
Interactive rendering of a huge dataset with data re-aggregation depending on zoom level
Here you can download the demo shown during the talk. The
demo is packed with anaconda-project and can be easily
started right after downloading. It automatically creates an environment with all required dependencies.
🐍️ Typed Settings Library
During one of the Lightning Talks, Stefan Scherfke presented his Typed Settings library. As the name implies, it is a library that provides a type-safe way to load settings from a configuration file. The following snippet shows a basic configuration definition and how to load it from a TOML file.
# pypirc.toml
[distutils.repos.pypi]
repository = "https://upload.pypi.org/legacy/"
username = "test"
[distutils.repos.test]
repository = "https://test.pypi.org/legacy/"
username = "test"
import sys
from typing import Dict
import typed_settings as ts
@ts.settings
class RepoServer:
repository: str
username: str
password: str = ts.secret(default="")
@ts.settings
class Settings:
repos: Dict[str, RepoServer]
settings = ts.load(Settings, "distutils", ["pypirc.toml"])
REPO_NAME = sys.argv[1]
print(settings.repos[REPO_NAME])
# RepoServer(repository='https://upload.pypi.org/legacy/', username='test', password=***)
I haven’t yet established a solid opinion on the merits of this library. Currently, I mostly use pydantic and load my
settings from a YAML file. Still, at first glance, the library looks like an interesting and flexible tool playing
nicely with other libraries like click and already mentioned pydantic. The functionality which I liked most is
the reading of settings from multiple sources. So I’ll keep an eye on it.
🐍 Running CPython in Browser
Wouldn’t it be great to run a Python interpreter in the browser? Not a re-implementation written in JavaScript or forwarding commands typed in a terminal-like window to a server, but the same interpreter you use on a laptop or remote machine?
Well, that what Christian Heimes presented in his Python 3.11 in the Web Browser - A Journey talk. As you might have guessed, the key ingredient is Web Assembly (WASM). It is possible (with some effort and restricted functionality) to compile the CPython interpreter to WASM and run it in the browser.
I’ll not go into the details here, as my experience with WASM and compiling complex C projects is not that good. But try to follow this link to access Python’s REPL and see what it looks like! Also, if you feel confident, check out the official CPython repository to access WASM build configuration and try it on your own.
🐍 ⚛️ Monitoring Machine Learning Projects
Lina Weichbrodt in her talk What I learned from monitoring more than 30 Machine Learning Use Cases reflects on her experience with monitoring machine learning projects. The key observation is that model evaluation and model monitoring are, in general, different exercises. When we evaluate a model during development, we are interested in such quality metrics as precision, recall, F1-score, and so on to estimate how well it performs. However, when it is deployed, we want to ensure that end users get the best possible experience. Therefore, we need to detect any anomalies in the model’s output and do it as early as possible.
In order to achieve this goal, the speaker proposes to rely on the “four golden signals”:
- Latency: the time it takes to serve a request.
- Traffic: the total number of requests.
- Errors: the number of requests that fail.
- Saturation: the load of your network and servers.
The following Venn diagram illustrates the concept. The set of metrics used in production includes but is not limited to metrics we use during development to monitor the quality of a model. Also, the speaker highlights the importance of input and output quality assurance. For example, detecting if inference requests have too many missing fields, checking if data distribution shifted, and other easily understandable “common sense” indicators.
 A model deployed in production requires additional set of metrics to quickly detect inference failures
A model deployed in production requires additional set of metrics to quickly detect inference failures
In other words, we shift our focus from how well a model performs on data to how reliable it is when serving requests. This way, we can ensure that “customer’s pain points” are detected and addressed as early as possible.
🐍 ⚛️ Managed Dask Clusters
Dask is a Python library for distributed computing. In essence, it takes such well-known libraries
like scikit-learn, numpy, and pandas, and scales them up to run on datasets that don’t fit into memory.
In some sense, it is like Spark, but for Python ecosystem. (I know that this comparison could be too superfluous, but
judging from my experience, it is fair enough.)
In the Spark world, there are multiple solutions that provide access to managed clusters, like Databricks or AWS EMR. Sure enough, at the start, nothing similar existed for Dask, as the technology wasn’t mature enough. But not anymore! Coiled provides a cloud platform for running Dask computations on managed clusters, helping users to easily scale their pipelines up.
In Data Science at Scale with Dask Richard Pelgrim conducted a hands-on tutorial of how to use Dask on its own and with Coiled-managed clusters. (See the repo to go over presented notebooks.) For example, a small snippet below shows running Dask on a local machine.
import dask.dataframe as dd
from dask.distributed import Client, LocalCluster
cluster = LocalCluster()
client = Client(cluster)
ddf = dd.read_csv(...).mean().compute()
How to run this on a managed cluster? Easy! Just replace your LocalCluster with coiled.Cluster.
import coiled
cluster = coiled.Cluster(name="bigdata", n_workers=10)
... # the same code as above
Because of close collaboration between Dask and scikit-learn development teams, it is also possible
to easily scale up some of its algorithms.
import joblib
from sklearn.model_selection import GridSearchCV
grid_search = GridSearchCV(...)
with joblib.parallel_backend("dask"):
grid_search.fit(data, target)
You can sign up and access clusters dashboard via Coiled cloud platform. For people who signed up during conference, some amount of free credits was granted. As far as I know, these credits are refreshed periodically and should be available for any person who wants to try the platform out.
The platform doesn’t (yet?) provide any managed notebooks or additional tools, except a cluster monitoring dashboard. Still, it feels like a great choice for developers and teams who don’t want to orchestrate computational infrastructure on their own.
⚛️ Data Version Control
There is a widely known concept in compute science: garbage in, garbage out. Machine learning engineers know it very well. The codebase is well-tested, the model is carefully tuned. But instead of getting high quality forecast, it starts to output some nonsense.
 Machine learning ain't easy
Machine learning ain't easy
Why? The chances are high that something changed in the data. A new encoding scheme applied to a certain categorical feature, too much label noise introduced in the latest data update, or any other data perturbation that deteriorated model’s performance significantly.
Data preprocessing and data quality are crucial for a stable and well-performing machine learning pipelines. Therefore, it is not enough to only track the changes in the codebase. One must take care of data version control as well. In his tutorial Making MLOps Uncool Again David de la Iglesia Castro presented a framework developed by Iterative helping to manage your data and run CI/CD pipelines tailored to data-driven projects. One of the core tools in this framework is the DVC library. As the name suggests, it enables version control of your data. But not only. It also tracks models and performance metrics, so can be treated as an experiment management system.
The tutorial demonstrates Iterative’s solutions helping to enable a CI/CD for an example NLP project. It explains how to use DVC to track changes in your data, and how to set up a reproducible pipeline for collaborative work.
A nice thing about this tutorial is that it doesn’t require any local setup and can be followed completely in browser via GitHub Web Editor. You only need to fork the repository and follow the instructions.
🌍 Inclusive Open Source
Finally, there is one more talk that worth mentioning. Melissa Weber Mendonça presented Beyond the basics: Contributor experience, diversity and culture in open source projects where she discusses the ethics of open source projects and such important topics as diversity and inclusion. The talk was presented remotely, as this time PyCon was running in a hybrid mode, supporting both on site and remote participants and speakers. And that was one of the points highlighted by the speaker, as such mode of conference is a great opportunity to participate for speakers and attendees from around the globe.
But there is more. The speaker reminds us about how much work is done by the open source community, and it is not only submitting pull requests and reporting issues. It also includes plenty of “invisible work” that is not reflected via repository metrics. Like solving organizational questions, engaging new contributors, volunteering. These contributions also should be renowned and rewarded.
Nowadays, many open source communities have Codes of Conduct and other guidelines helping to make them inclusive. However, there are other things that need to be addressed. For example, the speaker highlights the importance of more proactive approach of including contributors from underrepresented groups and regions.
PyCon is a great example of collective effort, driven by volunteers, experts and Python professionals who share their knowledge and put lots of effort to run the conference. It was an amazing opportunity to be there and meet people from all over the world sharing their ideas and projects. We should strive to make Python community as inclusive as possible, as it would benefit everyone.
Conclusion
In this post, I only briefly covered the topics that draw my attention during the conference. So I highly recommend you to go over the program and see if you can find something that would be interesting for you.
The conference was a great experience, and after two years of remote work and online meetings, it was nice to attend it in person for a change. Lots of fruitful conversations and well-prepared presentations. Plenty for cool frameworks of all kinds, from infrastructure orchestration to data management. I’m looking forward for the next year and wish to see you there, my fellow Pythonistas! 🐍
References
- Demystifying Python’s Internals: Diving into CPython by implementing a pipe operator
- Python Developer’s Guide
- CPython Internals
- Fast Native Data Structures: C/C++ from Python
- On Blocks, Copies and Views: Updating Pandas’ Internals
- Honey, I Shrunk the Target Variable!
- Seeing the Needle and the Haystack: Single-Datapoint Selection for Billion-Point Datasets
- Typed Settings Documentation.
- What I learned from monitoring more than 30 Machine Learning Use Cases
- Making MLOps Uncool Again
- Beyond the basics: Contributor experience, diversity and culture in open source projects





